A liquid biopsy platform combining a high fundamental frequency QCM device with dynamic chemistry for detecting mutations in circulating DNA
Authors: A. Grammoustianou, G. Papadakis, M. Tabraue, J.J. Díaz-Mochon, R. Fernández, J.V. García, A. Arnau, E. Gizeli. AWSensors S.L., Institute of Molecular Biology and Biotechnology- FORTH, University of Crete, Destina Genomics S.L., Centro de Investigación e Innovación en Bioingeniería – Universidad Politécnica de Valencia
Event: 5th International Conference on Bio-sensing Technology, Riva del Garda, Italy (2017)
In the past decade, the analysis of circulating tumour DNA (ctDNA) in blood has been a major breakthrough; ctDNA has been proposed as a priceless source for cancer diagnostic, prognostic and treatment monitoring through a new methodology known as “Liquid Biopsy”. This study presents a novel diagnostic method for the acoustic detection of KRAS mutations in ctDNAs based on: (1) DNA analysis by “dynamic chemistry” that utilizes aldehyde modified nucleobases (SMART) and abasic peptide nucleic acids (DGL probes) capable for the errorfree detection of nucleic acids and their mutations; and, (2) a high fundamental frequency (100 MHz) acoustic wave microsensor (AWS HFF-QCM) that allows the accurate, inexpensive, label-free and real time monitoring of the “dynamic chemistry”. Surface-immobilized DGL probes on the AWS HFF-QCM device are used to detect ctDNAs of wild type and mutated KRAS variants. Upon hybridization of the DGL probe with its target ssDNA, a duplex is formed where biotin- tagged SMART bases can lock in front of the position under interrogation; streptavidin binding detected in a follow-up step confirms the presence of the SMART bases. The use of DGL probes in combination with an isothermal DNA amplification step RPA) have allowed the sensitive and specific recognition of single mismatches in KRAS genes in less than 1 hour. This work presents a unique and novel technology that can emerge as a promising tool in the field of cancer diagnostics.
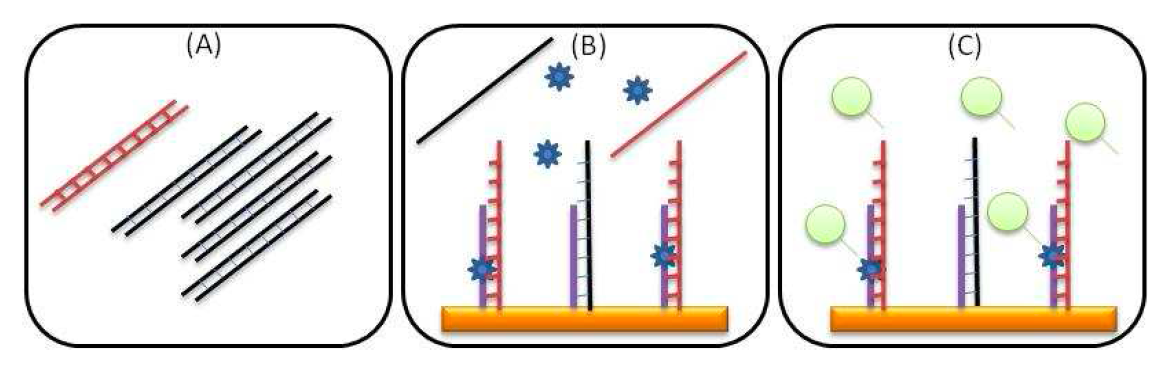
Schema for complete detection protocol:
(A) Extracted DNA containing mutant (red) and wild type DNA fragments (black) are enzymatically amplified.
(B) Denatured amplicons are hybridized on surface immobilized DGL probes. Chemical locking of a specific tagged SMART base takes place only in the appropriate position.
(C) Incorporated SMART bases are recognized by streptavidin and monitored in real-time during an acoustic measurement.


